Treating DNA Repair-deficient Breast Cancers
About 5 to 10 percent of breast cancers are hereditary, of which 20 to 25 percent bear alterations in the genes BRCA1 and BRCA2. These genes code for proteins in one of the DNA repair pathways, the homologous recombination (HR) pathway. Germline mutations (or inherited mutations) in these genes are widely recognized as conferring an increased lifetime risk for breast and ovarian cancers.
Defects in the HR pathway in human cells lead to DNA repair deficiencies, accumulation of DNA damage and, sometimes, cancer. When the HR pathway is defective, cells often use an alternate DNA repair pathway, called the base excision repair (BER) pathway, to fix their DNA. Normal cells do not replicate as often as cancer cells, so a class of cancer therapeutics called PARP inhibitors exploit this vulnerability of cancers. These therapeutics inhibit a key BER protein, poly ADP ribose polymerase (or PARP), thus preventing the cancer cells from repairing their DNA and forcing them to die. This mechanism of therapeutic targeting is referred to as synthetic lethality.
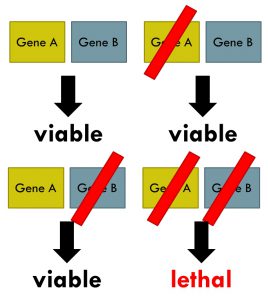
Synthetic lethality arises when a combination of deficiencies in the expression of two or more genes leads to cell death, whereas a deficiency in only one of these genes does not. Wikipedia image.
The U.S. Food and Drug Administration has approved three PARP inhibitors so far, olaparib (Lynparza) and rucaparib (Rubraca) to treat certain women who have ovarian cancer with BRCA gene mutations, and niraparib (Zejula) to treat both BRCA mutation-positive and BRCA mutation-negative ovarian cancers.
Several clinical trials are underway in which PARP inhibitors are being tested in breast cancers, mostly triple-negative breast cancers, because they often harbor BRCA mutations and DNA repair deficiencies. For example, in the phase III trial EMBRACA, the investigational PARP1/2 inhibitor talazoparib is being tested in breast cancer patients with BRCA mutations. Emerging studies show that the benefit of PARP inhibitors could extend beyond breast cancers with germline BRCA mutations.
PARP inhibitors may be effective against breast cancers with alterations in DNA repair genes other than BRCA
In a study published this week in the AACR journal Clinical Cancer Research, researchers from The University of Texas MD Anderson Cancer Center generated patient-derived xenografts (PDX; tumor models derived directly from patient tumors and grown in mice), to study different therapeutic options for residual triple-negative breast cancer after neoadjuvant chemotherapy.
Talazoparib caused regression of five of the 13 PDXs in which the drug was tested. Importantly, four of these five PDXs did not harbor germline BRCA1/2 mutations, but had somatic alterations in the genes involved in DNA repair, including PTEN, ATM, and BRCA2.

Unlike germline mutations, which arise in germ cells and are passed on to future generations, somatic mutations are alterations that arise spontaneously in any cell in the body and can accumulate during a person’s lifetime, sometimes causing cancer. The theory that somatic mutations and other alterations in DNA repair pathways may play a role in breast cancer was asserted in another study recently.
Alterations in DNA repair genes other than germline mutations in BRCA contribute to “signature 3”
Studies on genome-wide analysis of breast cancer tumors have revealed a distinct mutational signature, signature 3, associated with deficiencies of HR (HRD), represented by biallelic inactivation of BRCA1 or BRCA2.
An extended characterization of signature 3, published last month in Nature Genetics, showed that a large subset of tumors that had no significant germline defects in the BRCA genes had a mutational landscape similar to that of tumors with HRD, a phenomenon referred to as “BRCAness.” The study found that germline alterations in the gene PALB2, epigenetic silencing and somatic mutations in the gene RAD51C, and epigenetic silencing of BRCA1 were all associated with signature 3, and had the potential to abrogate HR function.
The authors note that these findings have therapeutic implications, as demonstrated by a preclinical study published in Molecular Cancer Therapeutics, in which RAD51C-deficient tumors were sensitive to the PARP inhibitor olaparib. In a phase II clinical trial called Talazoparib Beyond BRCA (TBB), the therapeutic is being tested in advanced breast cancer patients whose tumors have germline or somatic mutations in the HR pathway genes, including PTEN, PALB2, CHEK2, ATM, ATR, and RAD51C.
Our understanding of breast cancer and susceptibility continues to evolve rapidly. For example, genome-wide association analysis of more than 256,000 breast tumors and controls, reported recently in Nature, identified 65 new loci linked to overall breast cancer risk. These advances are critical for accelerating translational research, making way for improved breast cancer risk prediction and rational therapies.
The study published in Clinical Cancer Research was funded by the National Institutes of Health, the MD Anderson Cancer Center Moonshot Program, the Nellie B. Connally Breast Cancer Research Endowment, a Susan G. Komen Foundation for the Cure grant, The Cancer Prevention and Research Institute of Texas, the Barr funds, and a Stand Up To Cancer (SU2C) Dream Team Translational Research Grant (the AACR is the Scientific Partner of SU2C).



