Setting the Stage for New Cancer Therapeutics Through Basic Cancer Research
Basic cancer research provides the foundation for major breakthroughs in cancer treatment and discovery. To develop novel and effective therapeutics to treat a disease, we must first have an understanding of the molecular basis for that disease.
One of the many ways in which the American Association for Cancer Research supports basic cancer research is through its Basic Cancer Research Fellowship program. This one-year fellowship, awarded to several junior researchers annually, has provided early-career investigators with the necessary resources to pursue fundamental scientific questions since 1996.
Hugh Gannon, PhD, a recipient of the 2013 Anna D. Barker Fellowship in Basic Cancer Research, exemplifies the success of this program. While pursuing his doctorate at the University of Massachusetts Medical School, Gannon developed a strong interest in p53 biology as it related to cancer.
P53 is a “tumor suppressor” protein responsible for regulating cell division. Tumor suppressors like p53 are credited for preventing tumor growth by carefully modulating the process that can lead to tumor formation—excessive and uncontrolled cell division. However, often these proteins become damaged or mutated, resulting in protein dysfunction and the release of their control over cell division. Indeed, inactivation of p53 is observed in the majority of human cancers. As such, p53 is a heavily studied protein—but while many of its functions have been uncovered in relation to its role in disease, much remains to be discovered.
“Early in graduate school, I wanted to gain experience working with mouse models to study cancer,” says Gannon. “I was particularly drawn to p53-knockout mice due to their strong phenotypes in development and tumorigenesis.” However, Gannon recognized that the fundamental mechanisms governing the influence of p53 on tumor initiation and growth was still largely unknown.
After receiving his doctorate in 2012, Gannon began to explore these mechanisms in his postdoctoral work at the Dana-Farber Cancer Institute under the mentorship of Matthew Meyerson, MD, PhD. “As I was transitioning from graduate school to my postdoc, a number of papers were published that suggested the canonical pathways downstream of activated p53 (cell cycle arrest, apoptosis, and senescence) were dispensable for p53-dependent tumor suppression,” Gannon explains. “This sparked my interest in p53’s lesser-studied role in regulating oxidative stress in cancer.”
This was a new take on understanding the function of p53 in cancer. While p53-related therapies have mostly focused on reactivating wild-type p53 function, Gannon’s current work funded by his fellowship is seeking to exploit a pro-survival function of p53 that may be missing in cancers with disrupted p53 function. He is taking a large-scale approach, using expansive genomic datasets to identify potential therapeutic targets in cancer. These datasets include a comprehensive library of genomically characterized cancer cell lines (termed “Project Achilles”) used to screen for vulnerabilities within the genome, as well as the Cancer Cell Encyclopedia (CCLE), developed at the Broad Institute to characterize the molecular profiles of over 1,000 cancer cell lines.
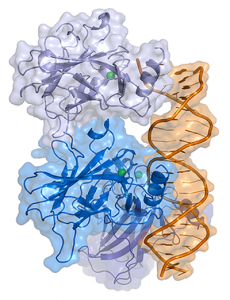
Representation of a complex between DNA and the protein p53. Image by Thomas Splettstoesser, licensed under CC BY-SA 3.0 via Wikimedia.
From these studies, Gannon has learned that like other downstream functions of p53, its role in regulating oxidative stress depends on cell type, genetic context, and cell culture conditions. Likewise, he has found that even the type of p53 mutation (such as missense mutations or deletions) can impact the cellular response to oxidative stress, with different types of mutations eliciting different responses. Of these observations, he says, ”We believe working with large-scale datasets will allow us to group and compare these different features (cell type, similar genetic contexts, types of p53 mutation) in a way that has not been done before due to the limited number of characterized cancer cell lines.”
What is unique about Gannon’s funded project is the ease with which it transitions between basic and translational research. Much of this transition can be attributed to the advances in genomics and sequencing, often employed to tease out these potentially important mutations that could serve as new targets for cancer therapeutics and benefit patients. “Sequencing studies can reveal potential drivers that can lead to targeted therapy development and identify mutations in rare patients that display exceptional responses in clinical trials,” Gannon says. “While this fast-paced translational work is compelling, I feel it is important at this time to further explore p53 function through basic research. The importance of p53 in cancer has been thoroughly established; once we fully understand why this is the case, effective treatment options in tumors with TP53 mutations will become apparent.”
Gannon is nearing the end of his fellowship term; however, his time as an AACR grantee has set the stage for a productive independent career by providing him with the necessary funding to make his innovative research ideas a reality. “This AACR grant has greatly impacted my training by allowing me to pursue a multidisciplinary project involving genomics, gene expression, and metabolism across hundreds of cancer samples and cell lines,” says Gannon. “These experiences have shaped the way I view and use large-scale datasets to independently generate my own hypotheses that can be potentially tested in the next phase of my career in cancer research.”
We look forward to seeing what is in store for the future of this bright young scientist.